Single-Cell Power
Biological systems are built from trillions of cells, but each cell can behave differently even within the same tissue. Traditional bulk sequencing averages signals across thousands of cells, masking the molecular fingerprints of rare but critical subpopulations. Single-cell sequencing overcomes this limitation by analyzing the transcriptome, genome, or epigenome of individual cells, revealing how cell heterogeneity drives health and disease. For example, in cancer, a small subset of cells carrying TP53 mutations may fuel tumor growth or therapy resistance, while in development, single-cell RNA sequencing can trace cell fate decisions as stem cells differentiate into specialized tissues. In immunology, profiling individual T and B cells highlights the remarkable diversity of immune responses to pathogens and vaccines. By identifying unique genetic or transcriptional states, single-cell technologies enable precision medicine approaches tailored to the specific cells driving disease, offering a molecular microscope into individuality that is transforming biology from basic mechanisms to clinical applications.
Spotlight on TP53
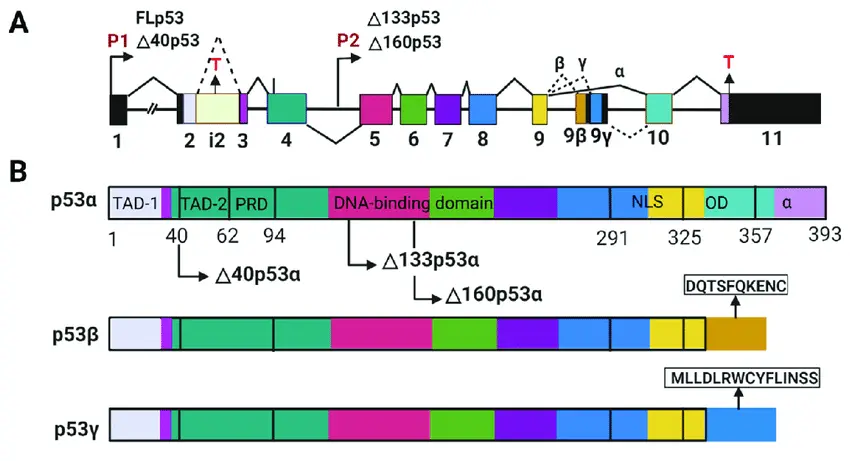
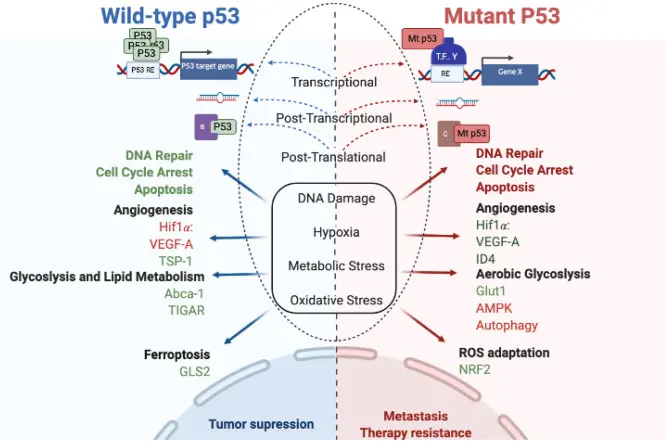
The TP53 gene, often called the “guardian of the genome,” safeguards our cells by regulating DNA repair, cell cycle arrest, and apoptosis. When functioning properly, TP53 prevents damaged cells from multiplying, acting as a powerful defense against cancer.
However, mutations in TP53 are among the most frequent genetic alterations in human cancers, and their effects can vary dramatically across individual cells. Single-cell sequencing reveals this hidden diversity, showing how some cells carry disruptive TP53 mutations while others remain unaffected. This imbalance fuels tumor evolution, therapy resistance, and relapse.
By mapping TP53 activity at single-cell resolution, researchers gain a deeper understanding of clonal diversity and disease progression. These insights pave the way for precision oncology, where treatments can be designed to target the specific subpopulations of cells that drive cancer growth.
Shifts in Cellular Dysfunction and Aging
Cells constantly face stress from environmental factors, DNA damage, or metabolic changes, and their ability to respond depends heavily on the TP53 gene. While TP53 normally acts as a safeguard, its activity can vary dramatically as cells age or accumulate mutations. Single-cell technologies allow researchers to examine these variations at unprecedented resolution, capturing how individual cells maintain or lose TP53 function.
In cancer, this heterogeneity is critical. A small subset of cells may acquire TP53 mutations that disable their protective mechanisms, allowing them to proliferate uncontrollably. These early mutant clones can expand, driving tumor formation, therapy resistance, and disease relapse even when the majority of neighboring cells remain healthy.
During aging, TP53 activity also changes in subtler ways. Declining TP53 function contributes to cellular senescence, the permanent withdrawal of cells from the cell cycle, which in turn affects tissue regeneration, organ function, and the overall aging process. By tracking these shifts at the single-cell level, scientists can map how disease and aging emerge from cellular diversity, revealing new targets for intervention and strategies to preserve tissue health over time.
Tracking Therapy Response
Cancer treatment outcomes are often determined by subtle differences between individual cells within a tumor. Single-cell technologies provide a window into these differences, allowing researchers to observe how each cell responds to therapy in real time. Cells with intact TP53 pathways may undergo apoptosis and be effectively eliminated, while neighboring cells with defective TP53 can survive, adapt, and eventually drive tumor relapse.
This cellular heterogeneity explains why treatments that appear effective in bulk analyses may fail in practice, as resistant subpopulations persist and expand. By monitoring these dynamics, scientists can identify which cells are likely to survive standard therapies and which therapeutic strategies are most likely to succeed.
Importantly, single-cell tracking enables the design of personalized interventions. By targeting the specific populations of cells that evade treatment whether through TP53 mutations or other protective mechanisms researchers can develop therapies tailored to the tumor’s unique cellular landscape. This precision approach holds the promise of improving treatment outcomes, reducing relapse, and ultimately transforming cancer care.
Decoding Cellular Resistance
Tumors are not uniform each cell can respond differently to therapy, and this variability often determines treatment success. Single-cell technologies allow researchers to decode these differences, revealing how some cells with intact TP53 pathways are eliminated, while others harbor mutations that let them survive and adapt. These resistant cells can expand over time, driving relapse and reducing overall treatment efficacy.
By mapping these responses at the single-cell level, scientists gain critical insights into the mechanisms behind therapy resistance. This knowledge enables the development of personalized treatment strategies that specifically target the subpopulations of cells most likely to survive, improving outcomes and reducing the risk of cancer recurrence.
Ultimately, decoding cellular resistance transforms our understanding of therapy dynamics, providing a roadmap for interventions that are both precise and effective, tailored to the unique cellular landscape of each patient’s tumor.